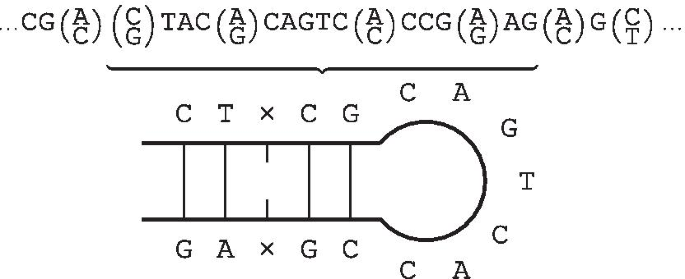
IUPACpal: efficient identification of inverted repeats in IUPAC-encoded DNA sequences | BMC Bioinformatics | Full Text
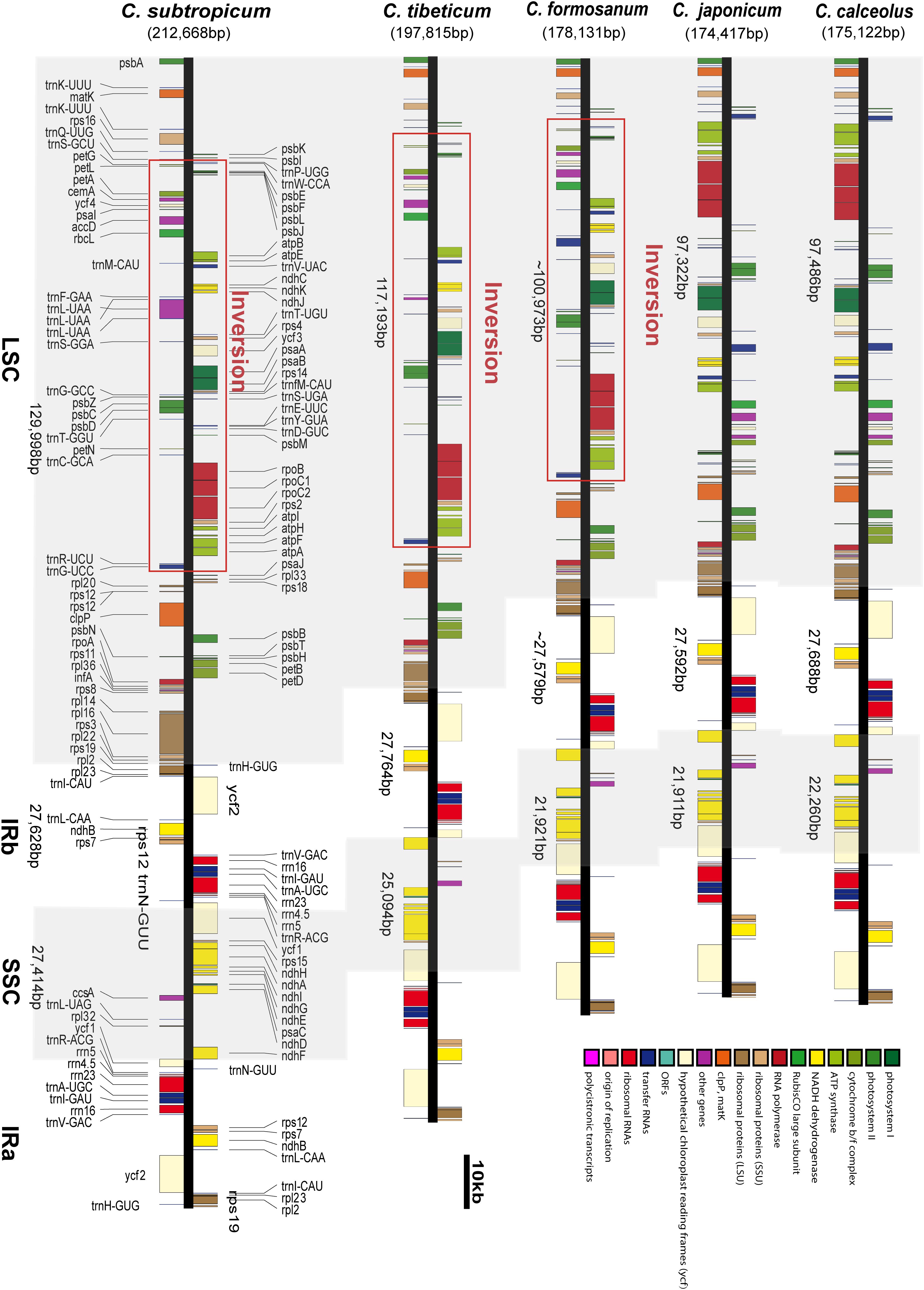
Frontiers | Chloroplast Genomes of Two Species of Cypripedium: Expanded Genome Size and Proliferation of AT-Biased Repeat Sequences | Plant Science
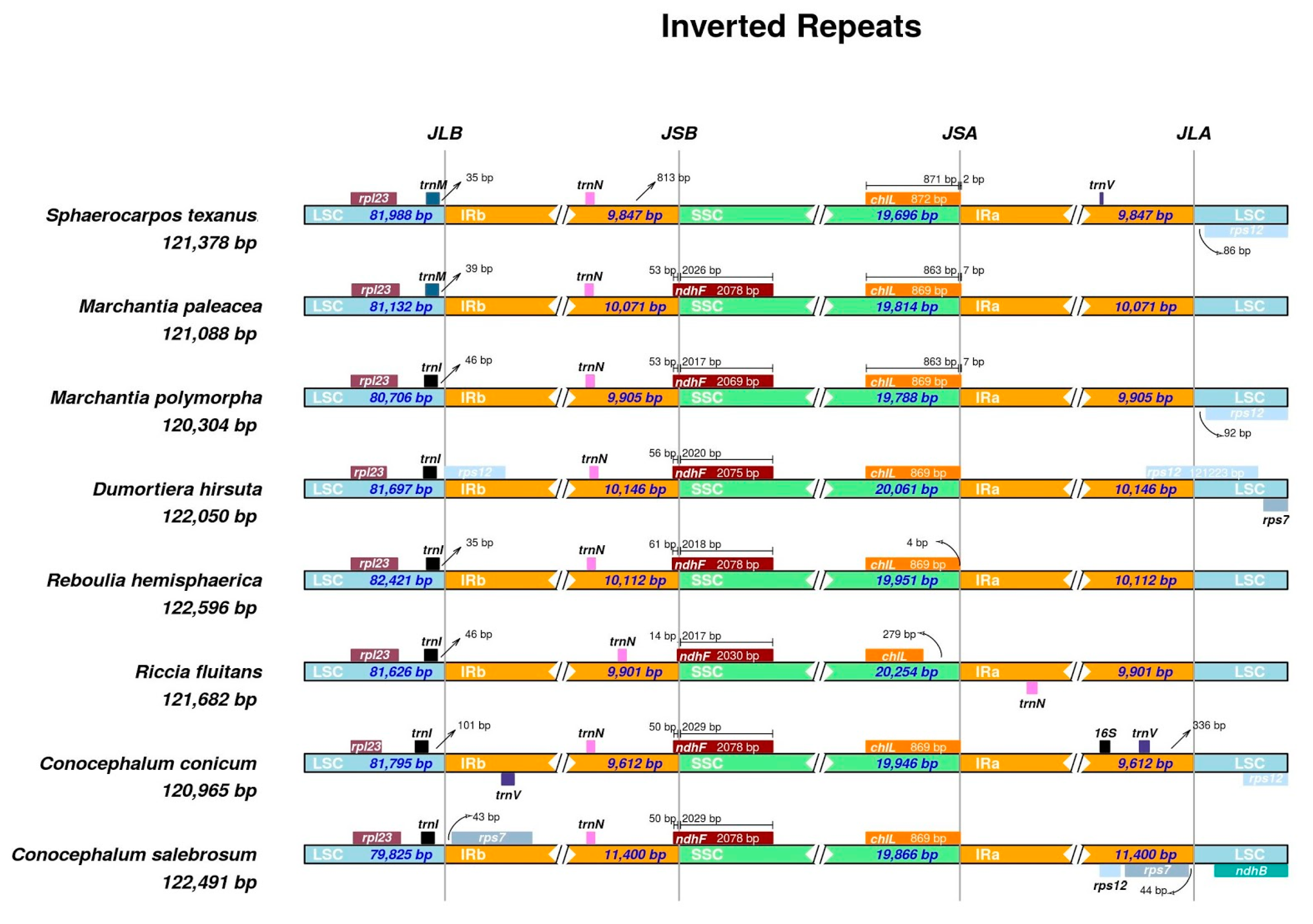
Genes | Free Full-Text | The Increase of Simple Sequence Repeats during Diversification of Marchantiidae, An Early Land Plant Lineage, Leads to the First Known Expansion of Inverted Repeats in the Evolutionarily-Stable
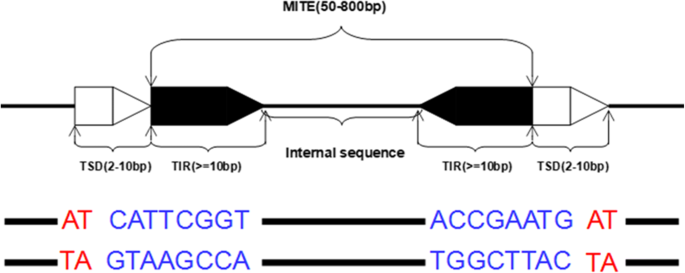
MiteFinderII: a novel tool to identify miniature inverted-repeat transposable elements hidden in eukaryotic genomes | BMC Medical Genomics | Full Text

Global analysis of inverted repeat sequences in human gene promoters reveals their non-random distribution and association with specific biological pathways - ScienceDirect

Inverted repeat motif as a potential origin of replication. Sequences... | Download Scientific Diagram

Expanded inverted repeat region with large scale inversion in the first complete plastid genome sequence of Plantago ovata | Scientific Reports

Insights on genome size evolution from a miniature inverted repeat transposon driving a satellite DNA - ScienceDirect